Football Table RL
Reinforcement Learning Basics
The football table employs on-line value iteration, namely Greedy-GQ[math]\displaystyle{ (\lambda) }[/math] and Approximate-Q[math]\displaystyle{ (\lambda) }[/math]. This page does not explain Reinforcement learning theory, it just touches on the usage and implementation of the provided library, which can be applied to any learning problem (libvfa located on the SVN). Too get a basic understanding of Reinforcement Learning i suggest reading the book by Sutton & Barto [1]. For a slightly more in-depth, hands-on, book on using RL with function approximation, i suggest the book by Lucian Busoniu (TU Delft) et al. [2], which is freely available as e-book from within the TU/e network. Unfortunately both books very different notations, here we use the notation from the former.
The update used for Greedy-GQ[math]\displaystyle{ (\lambda) }[/math] is:
[math]\displaystyle{ \vec{\theta}_{t+1} = \vec{\theta}_{t} + \alpha_t \left [\delta_t \vec{e}_t - \gamma(1-\lambda) (\vec{e}_t^{T} \vec{w}_t) \vec{\phi}(s_{t+1}, a^{*}_{t+1})\right ] }[/math]
[math]\displaystyle{
\vec{w}_{t+1} = \vec{w}_{t} + \beta \left [\delta_t \vec{\phi}_t - (\vec{\phi}^{T}_t \vec{w}_t)\vec{\phi}_t \right ]
}[/math]
With this is that and this is that. The update used for approximate-Q[math]\displaystyle{ (\lambda) }[/math] is:
[math]\displaystyle{ \vec{\theta}_{t+1} = \vec{\theta}_{t} + \alpha \delta_t \vec{e}_t }[/math]
From here on out this section assumes familiarity with reinforcement learning and discusses how the provided libvfa library works and is used. Note, this can be used on any project provided the user provides transition samples correctly.
Value function approximation
For this project it was chosen to use a Gaussian Radial Basis Function Network(RBFs) to approximate the Q-function, a short explanation on the method can be found here in [3]. Currently, this is the only method of function approximation available in the library, however other forms of linear parametric function approximation should be easy to incorporate (this requires to only change the membership function).
RBFs are a linear parametric function approximation, where the Q-function is estimated using:
[math]\displaystyle{ Q = \theta\cdot\phi(s,a)^{T} }[/math]
here [math]\displaystyle{ \phi(s,a)~(1 \times n) }[/math] denotes the so called feature vector, [math]\displaystyle{ \theta~(1 \times n) }[/math] denotes the parameter (or weight) vector and [math]\displaystyle{ n }[/math] denotes the number of nodes. The value of Q hence becomes a scalar, depended on the action and the state. During Reinforcement learning we are trying to estimate the vector [math]\displaystyle{ \theta }[/math]. A single member of a radial basis function is described like this (where the state is described by the vector [math]\displaystyle{ x }[/math]):
[math]\displaystyle{ \phi_i(x) = e^{\frac{||c_i-x||^2}{2(\sigma_i)^2}} }[/math]
The total feature vector has [math]\displaystyle{ n }[/math]-elements, each has a center [math]\displaystyle{ c_i\in S }[/math] and a width [math]\displaystyle{ \sigma_i }[/math]. A complete feature vector looks like this:
[math]\displaystyle{ \phi_i(x) = \left [\begin{array}{c}e^{\frac{||c_1-x||^2}{2(\sigma_1)^2}} \\ e{\frac{||c_2-x||^2}{2(\sigma_2)^2}} \\ ... \\ e{\frac{||c_n-x||^2}{2(\sigma_n)^2}}\end{array}\right ] }[/math]
Below, and example of a RBF network is shown. This particular example contains only three nodes and is 1-dimensional, with centers: [math]\displaystyle{ c = \left [\begin{array}{ccc} -x_{sat} & 0 & x_{sat}\end{array} \right ]^T }[/math]

Note from the above picture the difference between normalized and `regular` RBFs. Normalization can achieve slightly higher accuracy, however it does add to the computational demands. Normalization will make sure [math]\displaystyle{ \sum \phi(x)_i = 1 }[/math], where ([math]\displaystyle{ x = \left [ \begin{array}{cccc} x_1 & x_2 & ... & x_m \end{array}\right ] }[/math] for an [math]\displaystyle{ m }[/math]-dimensional problem). Naturally this leads to increasingly lower valued elements of [math]\displaystyle{ \phi(x) }[/math] for a higher [math]\displaystyle{ m }[/math]. This will effect the learning process, as values and differences become smaller, having the same effect as a low learning rate.
Formally, feature vector [math]\displaystyle{ \phi(s,a) }[/math], is depended on both the state and the action, creating one long vector that represents Q is its entirety (all combinations of actions and states). Basically allowing the feature vector have a different set of nodes, with different characteristics per action, changing the representation of the state-space per action.
However, to make things more simple and comprehensible libvfa is implemented slighty different(still correctly).
Instead of creating one long vector, we store [math]\displaystyle{ \theta }[/math] matrix of [math]\displaystyle{ (n_a \times n) }[/math] ([math]\displaystyle{ n_a }[/math] denotes the number of actions). This still holds the same information, however they are now stored in seperate rows, making indexation easier. On top of that, we use the same feature vector for each action (same node distribution and width across the states-space) yielding [math]\displaystyle{ \phi(s) }[/math] instead of [math]\displaystyle{ \phi(s,a) }[/math].
The latter is a simplification, and keeps compact. In some cases could be better however to tailor the feature-vector to the subset of the state-space in which the action can occur (which could also allow higher precision with a lower amount of nodes). I.e. in some parts of the state space some actions will never be available at all, whilst others are available. Hence the elements in the feature vector representing this area of the state space are useless for the non-available actions. Furthermore, it could be so that a different level of detail is required per action. This could be easily added to libvfa, investigating it's use would be quite interesting.
Sparse Calculations
One of the nice things of designing/defining a RBF network yourself, is that it allows you to use the knowledge of the network itself to speed things up. The main advantage can be obtained from addressing only nodes in the network that have a significant impact on the value we obtained; we are using the fact that it is a local function approximation. If we do this for every dimension, it quickly pays off, vastly reducing the size of vector we have to update, without compromising accuracy.
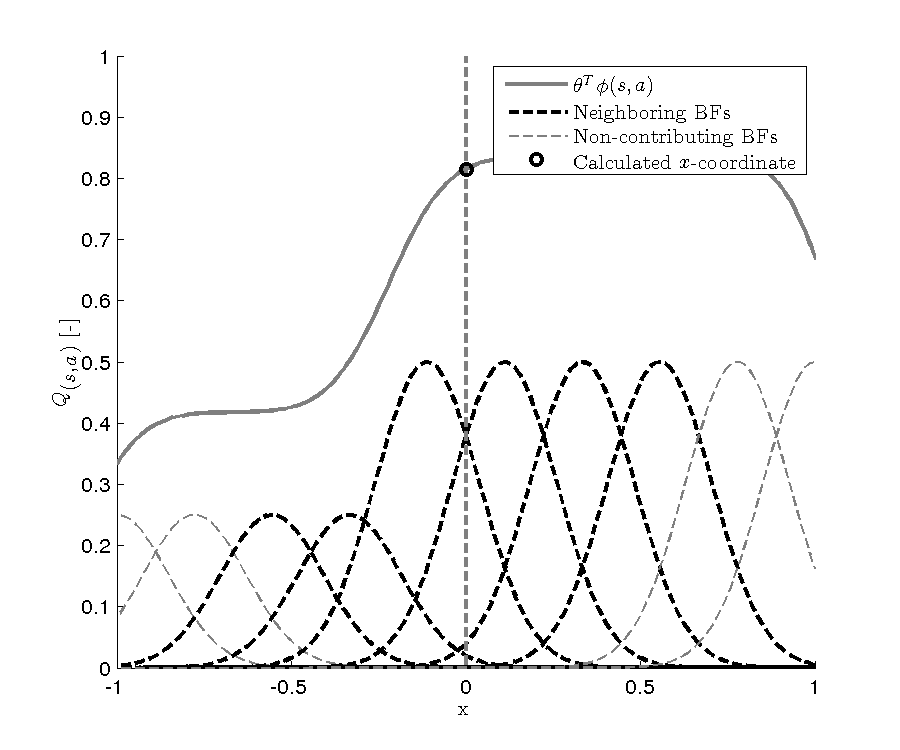
In libvfa, sparse calculations are executed whenever you call a function that ends with fast Fast. Also the traces are only implemented for the sparse calculations, so is the approximate Q-learning as a whole. In this library we first calculated what node lies closest to our current state, then depended on the width we include some nodes neighboring this closest center. In this library the width can be seen as a radius, a width of 2 will include the closest nodes and two nodes each side.Only these nodes are updated, so we never perform calculations on the whole vector.
When using eligibility traces, we need to make a small adjustment to the updates described in literature because in the trace vector, [math]\displaystyle{ \vec{e_t} }[/math], more states are `described'. This makes it more complex to create a sparse representation of this vector. So in order to avoid this we divide the update in two steps: a TD(0) step and a eligibility trace step: we seperate [math]\displaystyle{ \vec{e_t} }[/math], back into [math]\displaystyle{ \phi({s}) }[/math], [math]\displaystyle{ \phi({s_{t-1}}) }[/math], [math]\displaystyle{ \phi({s_{t-2}}) }[/math],.... , [math]\displaystyle{ \phi({s_{t-n}}) }[/math], and do updates on these states seperately. Doing the eligibility trace step this way requires a buffer of previous states. After a transition we first update the current state, [math]\displaystyle{ s }[/math], which is called the TD(0) update. Then this we assign the eligibility trace to the preceding states iteratively (see an example of the implementation here ). This relieves us having to store keep and update the full-sized [math]\displaystyle{ \vec{e_t} }[/math], whilst still achieving the same update. This is a lot faster, however if traces become really deep (not recommended), it might become better to just update the full problem more traditionally.
Library functions
In this section we explain how to use some of the important functions of libvfa. The best way too familiarize yourself is to read primitives_select.cpp. But here i present some basic explanation.
Initialize
Initialization is done using either the constructor or vfa::init.
int numberOfActions = 2;
int numberOfStates = 2;
double numberOfNodesPerState[2] = {3, 3};
double saturationPerState[2] = {1, 1};
double nodeWidthPerState[2] = {1, 1};
bool normalize = false;
bool varRate = false; //this does not work well
vfa EXAMPLE(numberOfActions, numberOfStates, numberOfNodesPerState, saturationPerState, nodeWidthPerState, normalize, varRate);
Or you can do this:
vfa EXAMPLE;
EXAMPLE.init(numberOfActions, numberOfStates, numberOfNodesPerState, saturationPerState, nodeWidthPerState, normalize, varRate);
The variables used for initialization in the previous example are explained below:
(int) numberOfActions- Total number of actions available to the decision maker (size only limited by memory).
(int) numberOfStates- Total number of state available to the decision maker (maximum currently is 8, expansion should be relatively simple (instructions in the code).
(double *)numberOfNodesPerState- Total number of nodes per state (above example code would yield ([math]\displaystyle{ 3 \times 3 }[/math]) nodes), equidistantly devided over the statespace.
(double *)saturationPerState- The (absolute) saturation per state, i.e. using 1 would yield nodescenters to be distributed between [math]\displaystyle{ -x_{sat}\leq c_i\leq x_{sat} }[/math]. (Nodes are always placed on the 'edge'([math]\displaystyle{ x_{sat} }[/math]) of the defined space
(double *)nodeWidthPerState- Width of the nodes, per state ([math]\displaystyle{ \sigma }[/math]). It is recommended to use (for state [math]\displaystyle{ i }[/math]):
[math]\displaystyle{ \sigma_i = \frac{x_{i,sat}}{n_{i,nodes}-1} }[/math].
This yields a width of half the distance in between the nodes. (bool) normalize- Indicate whether you want to normalize the RBFs, this can be useful when using a small amount of nodes over a large interval (see figure)
(bool) varRate- Indicate if you want to use a variable learning rate, based previous on activation of nodes in the past. Does not work very well and is overly complicated.
Getting Values
If we want a single Q-value, we can use this:
int action = 0;
double state[2] = {0,0};
int width = 2; //Here we determine the number of neighbours we use
double value = EXAMPLE.getValueFast(action, state, width)
int width here describes how many surounding neigbouring nodes are use for approximation (per side, symmetrically). If we want the values of a couple of particalur actions
int nActions = 3; int actions[3] = {0,2,3};
double state[2] = {0,0};
int width = 2; //Here we determine the number of neighbours we use
fvec values = EXAMPLE.getValuesFast(state, actions, nActions, width)
If we want all, we can use getAllValuesFast. Using the last two we get a vector, which allows us to easily find the maximum:
uword maxQIndex ; //uword is simular to int
double maxQ = values.max( maxQIndex); //Gives the maximum value and writes the index(in fvec values) to maxQIndex
Performing updates
To perform an in Q-learning update we need the [math]\displaystyle{ s,~a,~s_{t+1},~r_{t+1} }[/math]. Also currently we provide all the learning parameters as well ([math]\displaystyle{ \alpha, \beta, \gamma, \lambda, \epsilon }[/math]). You can choose either approximate-Q or Greedy-GQ to perform the update. For Greedy-GQ we use:
VFA.updateGQReallyFast(double * currentState, double * nextState, int action, double reward, double alpha, double beta, double gamma, double lambda , int width);
For approximate Q we use:
VFA.updateGQReallyFast(double * currentState, double * nextState, int action, double reward, double alpha, double gamma , int width);
Doing eligibility traces
Eligibility traces are done separately after a normal update, to perform such an update use:
VFA.updateGQLambda(double * traceState, int traceAction, double alpha,double beta,double gamma,double lambda , int width, int depth);
Here the int depth states how many steps back we are from the current state ([math]\displaystyle{ s_{t-depth} }[/math]). Similarly for approximate-Q we can use:
VFA.updateTDLambda(double * traceState, int traceAction, double alpha,double gamma,double lambda , int width, int depth);
So to perform an update on all previous states within the episode you need to loop back using a buffer. Here we have some example code, where we use phiBuf to break off the trace once an sub-optimal action was taken (if useWatkins is true).
for(int i = 1; i<traceDepth_; i++){
- traceIndex = bufIndex - i;
- /*Same episode && either (no Watkins || Watkins and we took an 'optimal' action)*/
- if(eBuf(traceIndex) == eBuf(bufIndex) && (!useWatkins || useWatkins && phiBuf(traceIndex+1)!=0)){ //Do not break
- }else{ break;}
- for(int j = 0; j<nStates; j++){stateTrace[j] = csBuf(traceIndex, i);}
- if(useGQ_){
- VFA.updateGQLambda(stateTrace, aBuf(traceIndex), alpha_, beta_, gamma_, lambda_, epsilon_, 2, i);
- }else if(useApproxQ_){
- VFA.updateTDLambda(stateTrace, aBuf(traceIndex), alpha_,gamma_, lambda_, epsilon_, 2, i);
- }
}
- ↑ Reinforcement Learning: an introduction, Sutton & Barto, http://webdocs.cs.ualberta.ca/~sutton/book/the-book.html
- ↑ Reinforcement Learning and Dynamic Programming using Function Approximation, Lucian Busoniu , Robert Babuska , Bart De Schutter & Damien Ernst, 2010, http://www.crcnetbase.com/isbn/9781439821091
- ↑ Comparison of CMACs and radial basis functions for local function approximators in reinforcement learning, R.M. Kretchmar && C. W. Anderson, http://www.cs.colostate.edu/pubserv/pubs/Kretchmar-anderson-res-rl-matt-icnn97.pdf